Meet the nano-sized rabbit made of DNA
Scientists have developed a new automated method to fold DNA strands into complex 3D structures. The new results will help pave the way for DNA nanotechnology in medicine over the next 10 years.
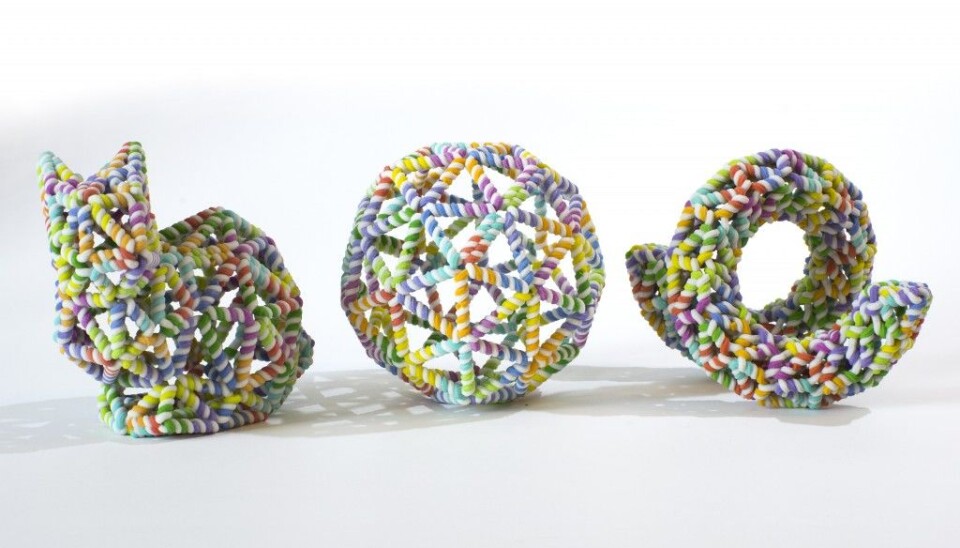
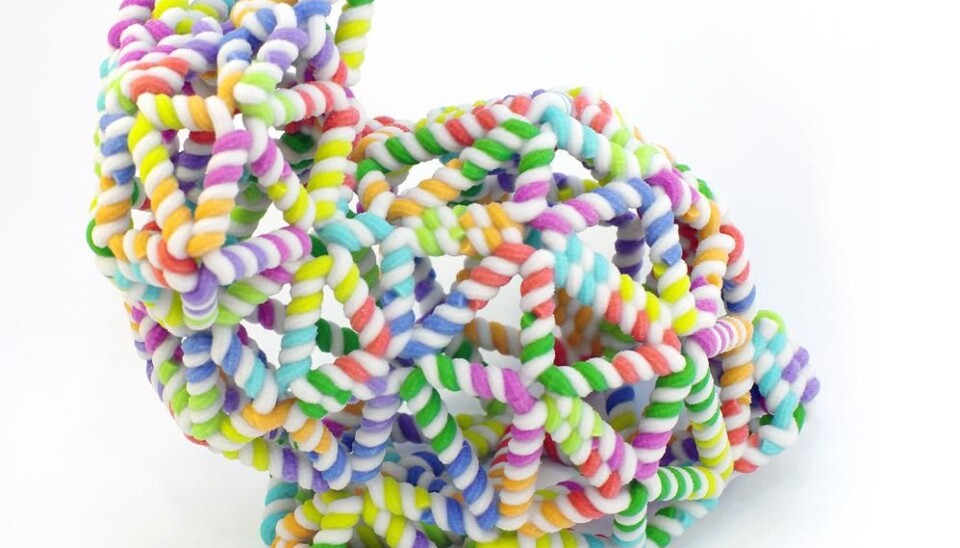
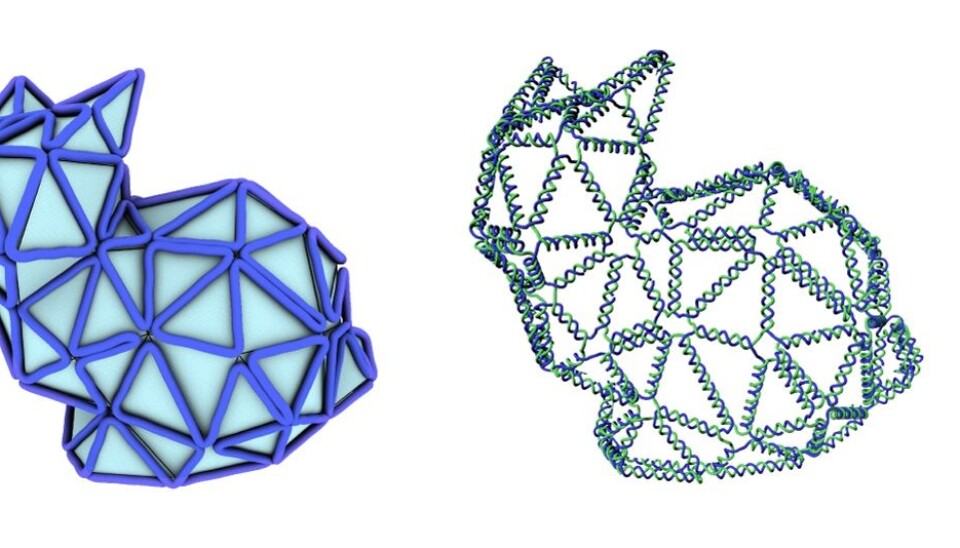
Meet the nano-sized rabbit, made entirely of DNA strands. It is designed using a new method that simplifies the existing design process of these tiny structures and brings us a step closer to targeted drug delivery and other medical applications using similar DNA structures.
“We’ve developed a new slim lined method to fold DNA strands into more complex structures than previous attempts,” says Björn Högberg, a senior lecturer at the Karolinska Institute, Sweden, and head of the research group behind the new study.
“We use the same principles as 3D printing to design the DNA structure, and most importantly we have shown that these designs work in real life, by developing some examples in the laboratory,” he says.
These complex structures are less than 50 nanometres wide and have been produced as part of a new study published this week in the journal Nature.
What is DNA folding?
DNA origami has been around for nearly 10 years now, and since then we have been focussed on perfecting the design principles of the DNA nanostructures.
Also called DNA-origami, it describes the process of encouraging DNA to fold itself into 2D and 3D structures.
It uses the natural properties of DNA double helix, whereby each of the 4 bases (A, T, C, and G) is attracted to another base to create a complimentary pair. By attaching lots of short DNA strands at certain points along a longer strand of DNA, the scientists help this long DNA strand to fold at precise points and ultimately form into 2D and 3D structures.
According to Ebbe Andersen, assistant professor in the Interdisciplinary Nanoscience Center, at Aarhus University in Denmark, DNA origami has been around for a while, but the new methods developed by Högberg and his team, provides a general approach, opening it up to people outside the field of DNA research.
Andersen was not involved with the new research, but he also studies DNA folding techniques. He created the first 3D origami box out of DNA in a study published in the journal Nature in 2009.
“In previous software you have to build up a design helix by helix, but this group use a different approach, concentrating instead on the shape you want to achieve, and then the software figures out the details for you,” he says.
This is echoed by Mark Bathe, associate professor of biological engineering at MIT, in the US, who also studies DNA nanotechnology and was not involved with the new study.
"This work takes an important step towards fully automating DNA-based materials design at the nanometre-scale,” say Bathe, who describes the new design approach as very flexible in its use of computer graphic techniques.
A new method to simulate DNA-nanostructure design
The new method is based around a more automated design process.
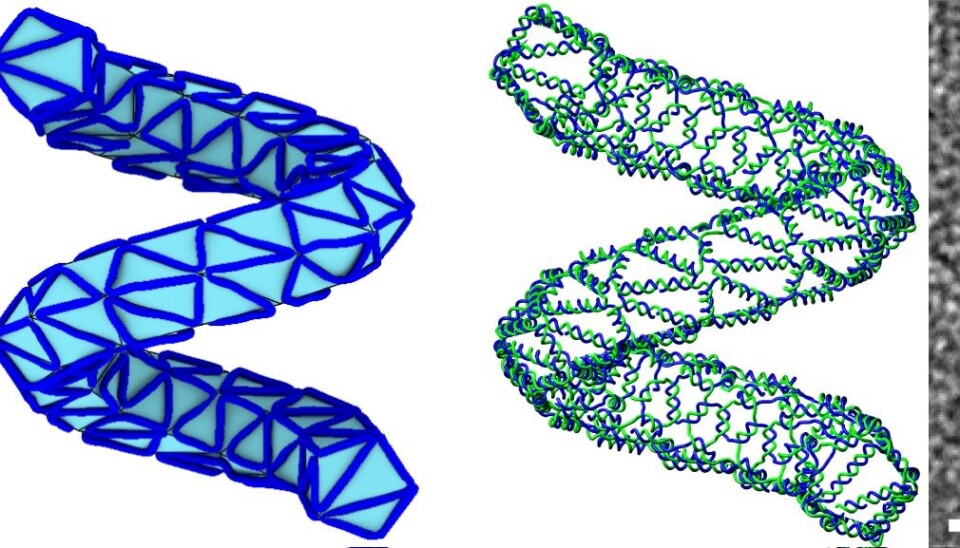
First, they design the structure on the computer, for example of a rabbit, or a ball, using a series of polygons to create a mesh. Then they use a newly developed algorithm to tell them how to organise the DNA strands along the edges of this mesh, to achieve the final desired shape in DNA.
“We take the paradigm of 3D printing to construct a method to go from a mesh designed in the computer, and use a new algorithm to simulate a design in DNA that looks like this mesh,” says Högberg.
“The trick is to make more complicated structures, whilst using as little DNA as possible, which is where the algorithm really helps,” he says.
From the computer to the laboratory
Designing shapes on a computer is one thing, but the scientists also needed to be able to actually construct these nanostructures in real life, back in the laboratory.
“Perhaps the most important part of our research is that we have implemented this in reality,” says Högberg.
“It’s not just that we have a useful computer algorithm, but we have a real method to produce more complex DNA structures. And we show this by successfully folding a bunch of DNA, very precisely, into the desired shapes,” he says.
So far, they have created a nano-sized rabbit made out of DNA -- the so-called Stanford bunny commonly used to demonstrate 3D printing techniques -- amongst other interesting shapes, such as a ball, a bottle, and even something that closely resembles a person waving.
Medical applications now and in the future
Behind the fun side of creating microscopic rabbits out of DNA, the research has significant real-world applications.
Andersen expects the new method to help other researchers to develop the tiny structures needed for targeted drug delivery in, for example, cancer treatments, as the new structures function better inside the human body than previous attempts.
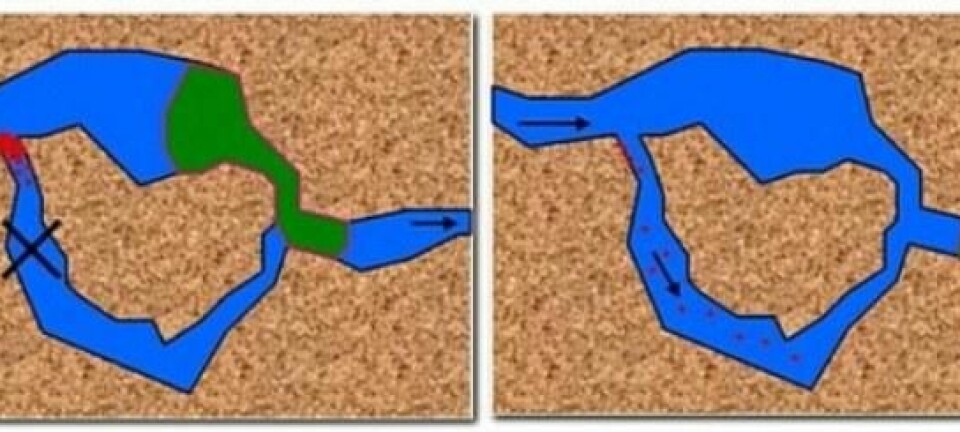
“In previous designs the helices are packed tightly together, and so they can only function in a liquid solution that contains a lot of magnesium, which counteracts repulsion between these negatively charged helices. But this new design has more space between the helices, so you don’t need this extra magnesium. This makes it function better in a biological environment, such as inside the human body where magnesium levels are lower” says Andersen.
The new design method is also helping Högberg and his team to study how cells communicate in the body, and to create structures for medical treatments.
“The cells in our bodies communicate by adding proteins onto their surfaces, and the neighbouring cells have receptors to detect these proteins,” says Högberg.
“You can think of it like brail reading for the blind, but instead of reading by dots, the cells read by these nanoscale protein clusters on neighbouring cells,” he says.
“We are right now developing our own artificial clusters, using similar DNA-nanostructures presented in this research, and these structures will be a guide for developing future research, including drug delivery systems, possibly in the next 5 to 10 years,” says Högberg.






.gif)


