Scientists can now discover global epidemics before they start
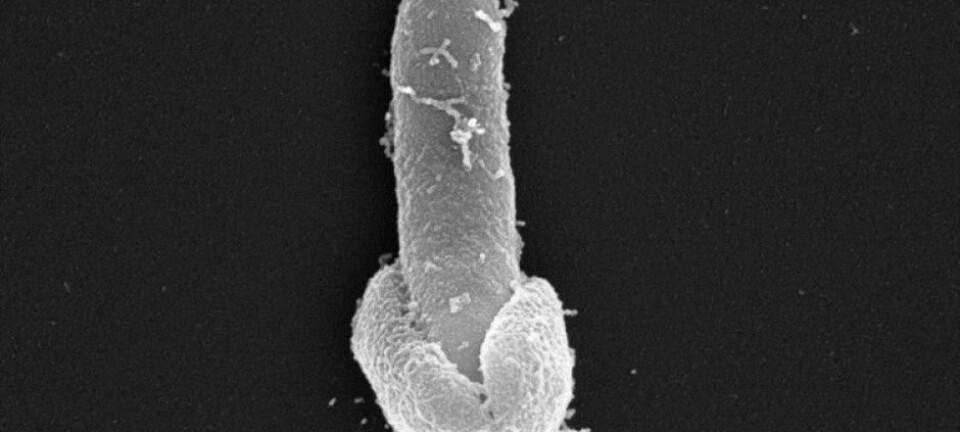
Scientists studying ‘flesh-eating’ bacteria discover how bacterial epidemics start. The results offer hope for new diagnostic tests to stop future epidemics in their tracks.
Scientists have sequenced the entire genome of almost 5,000 strains of bacteria known as group A Streptococcus, which is responsible for 600 million infections worldwide and can cause a nasty ‘flesh-eating’ disease in severe cases.
They have identified a specific region in the bacterium genome, where genetic mutations produce toxic proteins that are particularly virulent and harmful to humans.
Until now scientists did not know what triggered the precise molecular changes in the bacteria’s genome that leads to such epidemics or how particular types of bacteria, like group A Streptococcus, become so virulent in the first place.
“We can now pin-point two specific nucleotide changes and the exact genetic events that increase the production of certain harmful toxins, causing these bacteria to become so virulent and spread around the world as a global epidemic” says Professor Karl Kristinsson, head of Clinical Microbiology Department at the National University Hospital of Iceland and co-author on the study.
The scientists behind the study expect that other researchers will now be able to design rapid diagnostic tests to identify these genetic events and reduce or halt future epidemics before they spread.
Harmful toxins created in the genome make a virulent strain
For decades, scientists have wondered how particular strains of bacteria become so virulent and spread around the world so quickly as an epidemic.
In the new study published this week in the Journal of Clinical Investigation, the scientists have sequenced strains of group A Streptococcus bacteria (otherwise known as group A strep) to pin-point genetic changes in two types of the bacteria, called M1 and M89.
“Our hypothesis when we began the research was that there were going to be genetic changes in group A strep bacteria that contributed to epidemics,” says the senior scientist on the study, James Musser, professor of pathology and genomic medicine at Houston Methodist, USA, in a video that accompanies the new study.
“We sequenced the genome of thousands of strains, precisely defining every base pair in the strain. Fortunately, our hypothesis turned out to be correct. There are changes in the germ, the pathogen, that contribute to the epidemics,” he says in the video.
They saw changes in specific genes in a region of the genome known as the regulatory region. This area regulates the production of two harmful toxins--proteins called NAD+-glycohydrolase and streptolysin O.
“Every gene has what is called a regulatory region, it is involved in how genes are transcribed and how proteins are made. Changes that have occurred with group A strep involve changes in the regulatory region and that results in increased production of [these two toxins],” says Musser.
“So it’s really not so much mutations within the gene itself, but they’re actually in the regulatory part of the gene,” he says.
This precise genetic trigger makes these strains of group A strep bacteria virulent enough to spread as an epidemic.
Video: Houston Methodist.
An international collaboration to pin-point the cause of epidemics
The new study expands upon a previous study published in the journal PNAS in 2014, where the same team of scientists sequenced the entire genome of 3600 strains of one type of strep bacteria known as type M1.
“This all started a few years ago, as a collaboration between Musser’s group and others in the US and research institutes in Finland, Iceland, Denmark, Sweden, and Canada,” says Jaana Vuopio, professor of bacteriology at the University of Turku, Finland. She is a co-author on both studies.
“In the 2014 study we looked at one strain type, M1, which started to evolve and change its composition in the 1980s, and it was at that time that the disease changed and spread around the world as an epidemic,” she says.
They identified the molecular events, certain genetic changes, which created the two particularly toxic proteins. And they saw that these genetic changes occurred around 1983--right before the global M1 epidemic.
But they still did not know precisely what mechanism had triggered these genetic events in the first place and caused the spread of this single type of particularly virulent bacterium. They performed new genomic experiments and biological virulence assays to investigate this change.
“In addition, we looked at an additional strain of group A strep bacteria called M89, which is also spreading, and becoming more frequent in many countries, including Finland and US,” says Jaana.
“We repeated the same type of whole genome analysis as we did in the 2014 paper for a large group of these new M89 strains, and we found the exact same genetic mechanisms that we had previously observed in the evolution of M1,” she says.
Stopping epidemics before they start
According to all the scientists involved in the study, they are now able to quickly pin-point the genetic changes in bacteria that could signal an epidemic.
They expect other research groups to turn their findings into diagnostic tests to identify the changes and either halt an epidemic in its tracks or at least to reduce its spread around the world, says Kristinsson.
“Developing these tests could be relatively fast now that we have provided the gene sequences. For example, it will be possible to design DNA probes for routine medical tests. Whether this will become a commercially produced product, only time will tell,” he says.
Scientific links
- A molecular trigger for intercontinental epidemics of group A Streptococcus. 2015. DOI:10.1172/JCI82478
- Evolutionary pathway to increased virulence and epidemic group A Streptococcus disease derived from 3,615 genome sequences. 2014. DOI: 10.1073/pnas.1403138111.